AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
Back to Blog
Clustering app cytoscape4/30/2023 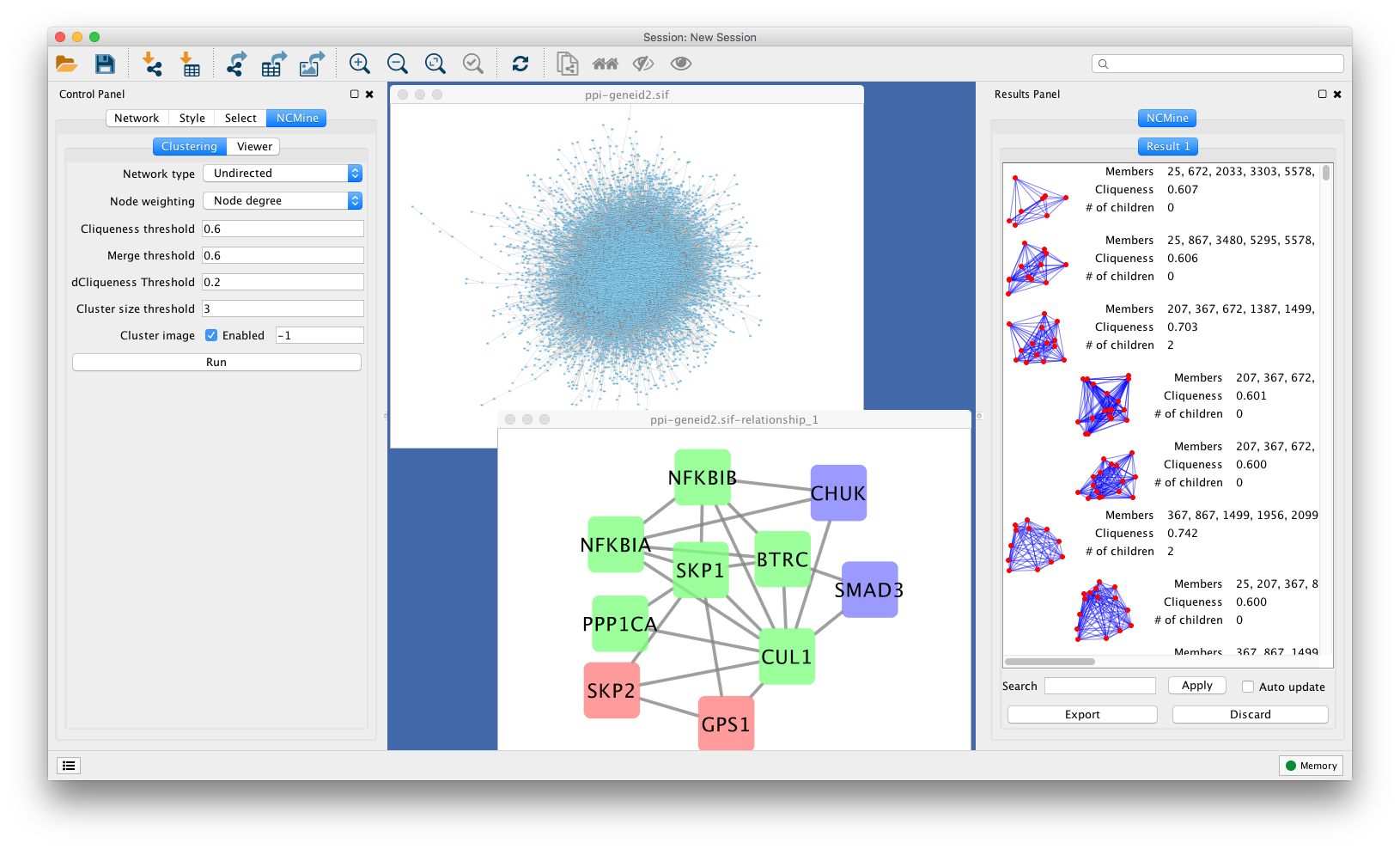
ClusterONE Cytoscape Plugin Finding overlapping protein complexes in PINs CytoCluster Cytoscape Plugin Detecting overlapping protein complexes in PINs ClusterViz Cytoscape Plugin Finding clusters in PINs using various clustering algorithms. The role proteins play within the cell is a highly efficient, pre- Table 3 Some popular tools for modularity analysis Tools Platform Description MCODE Cytoscape Plugin Finding densely connected regions based on topology in PINs. The selected key gene GNB4 is a potential biomarker to guide the immunotherapy of gastric cancer. qRT-PCR demonstrated that GNB4 expression in gastric cancer was notably higher in comparison with that in normal gastric mucosa, showing significant association with matrix TILs. 
The GSEA, TISDIB, and TIMER databases revealed that GNB4 is involved in various tumor signal pathways and immune and metabolic processes. Oncomine and GEPIA databases revealed that GNB4 expression in gastric cancer was obviously higher in comparison with that in normal gastric mucosa. Furthermore, 18 genes were screened as hub genes on the basis of the univariate Cox risk model of TCGA database (82 differential genes predicted poor GC survival). Overall, altogether 656 genes were upregulated with 5 genes being downregulated, which were matrix immune-related differential genes. Patients undergoing a great stromal score exhibited worse prognostic outcome, and cases having a low immune score had better prognosis. qRT-PCR was applied for exploring differential GNB4 expression between GC and normal gastric mucosa and investigating the relation of GNB4 with tumor-infiltrating lymphocytes (TILs). At last, we adopted TISDIB and TIMER databases for analyzing the association of guanine nucleotide binding protein subunit-4 (GNB4) between gastric cancer and tumor immune cells. 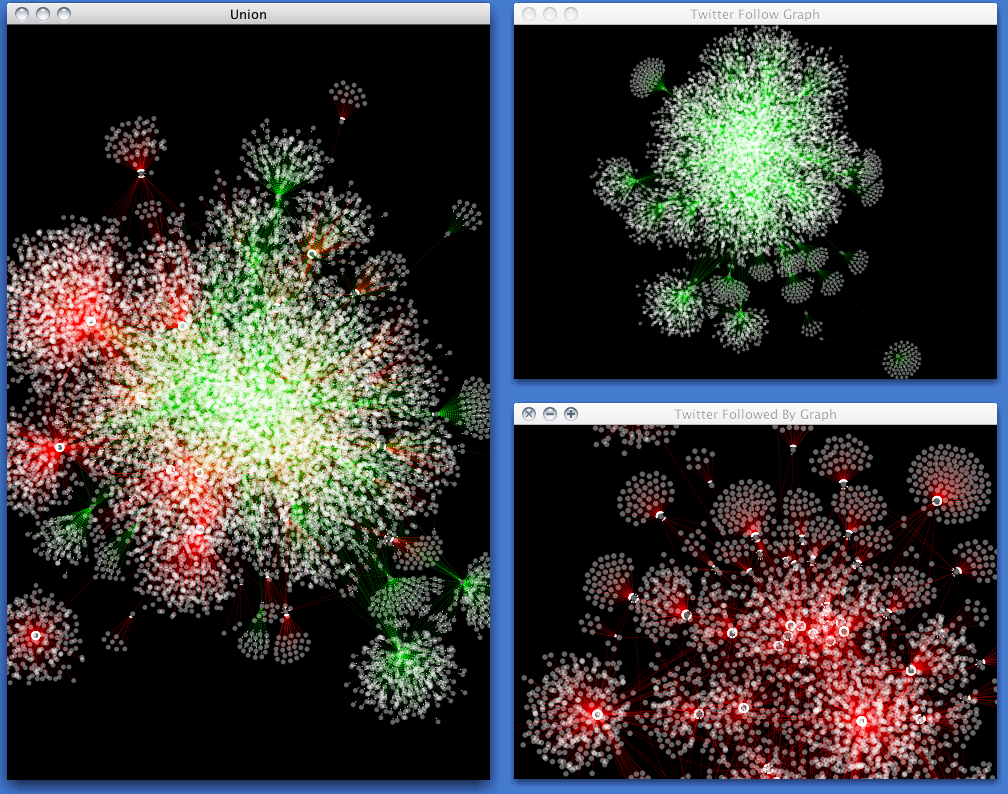
Oncomine and GEPIA databases were used, aiming to study the differences in key genes in healthy gastric mucosa and GC. The MCODE module of Cytoscape software was used to screen key genes. Besides, the STRING database was involved in order to detect the association among the proteins. This work employed Sangerbox to explore the differentially denoted genes (DEGs) related to stromal, immunity, and prognosis. 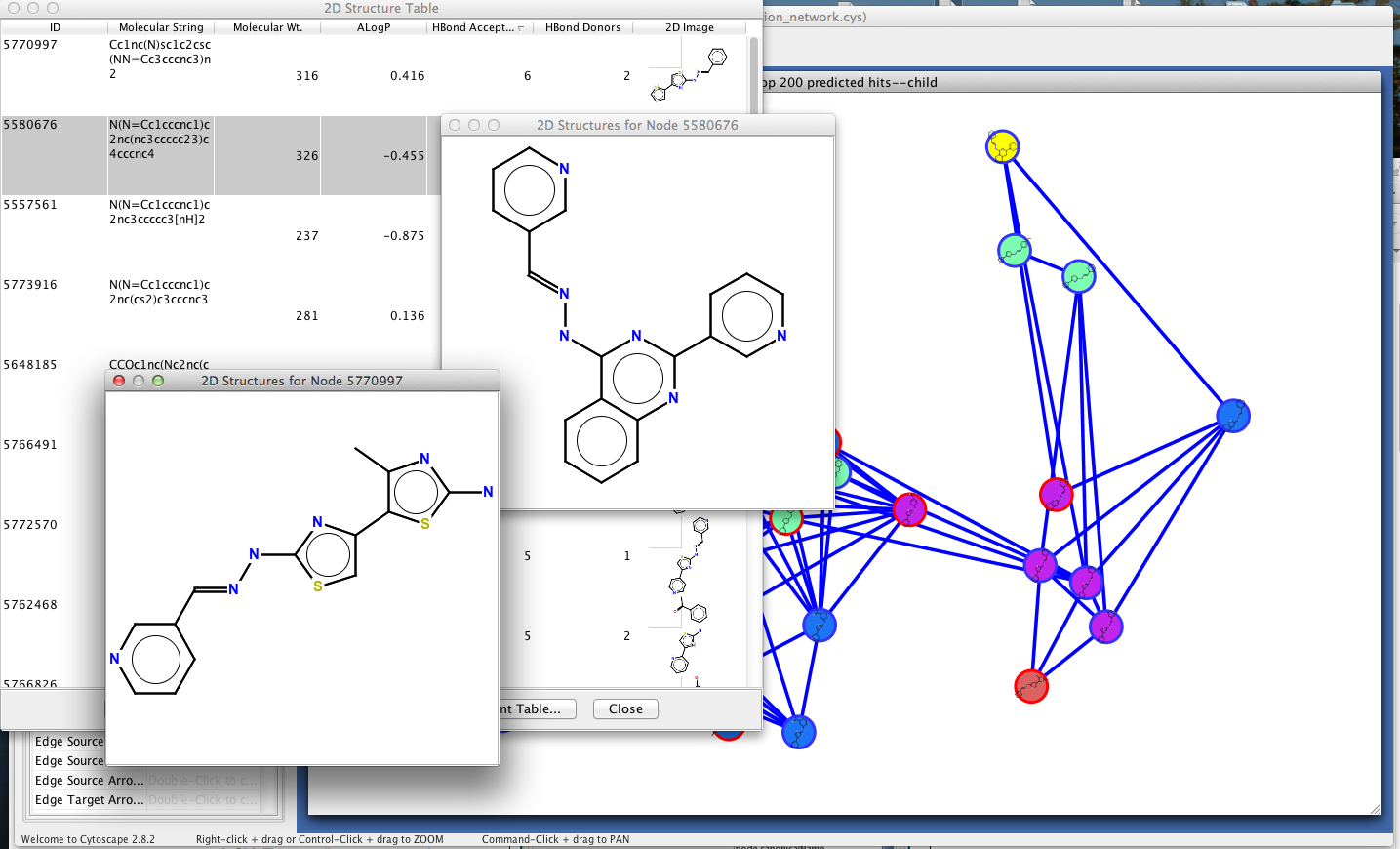
The ESTIMATE algorithm was adopted for evaluating the score of immune-infiltrating cells. Stromal and immune cells belong to two main nontumor components exerting a vital function in the tumor microenvironment.īased on TCGA database, this study downloaded clinical information and gene profiles of GC. Gastric cancer (GC) represents a universal malignant tumor of the digestive system. To illustrate usability of ClusterViz, we provided three examples with detailed steps from the important scientific articles, which show that our tool has helped several research teams do their research work on the mechanism of the biological networks. Due to adopting the abstract interface of algorithms in module of the clustering algorithms, more clustering algorithms can be included for the future use. Three commonly used clustering algorithms, FAG-EC, EAGLE and MCODE, are included in the current version. ClusterViz fascinates the comparison of the results of different algorithms to do further related analysis. According to the architecture, the implementation of ClusterViz is partitioned into three modules including interface of ClusterViz, clustering algorithms and visualization and export. In order to reduce complexity and enable extendibility for ClusterViz, we designed the architecture of ClusterViz based on the framework of Open Services Gateway Initiative. In this paper, ClusterViz, an APP of Cytoscape 3 for cluster analysis and visualization, has been developed. Furthermore, visualization of clustering results is crucial to uncover the structure of biological networks. Cluster analysis of biological networks is one of the most important approaches for identifying functional modules and predicting protein functions.
0 Comments
Read More
Leave a Reply. |
 RSS Feed
RSS Feed
